| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-09-30 22:30:25 UTC |
|---|
| Update Date | 2020-05-11 20:20:50 UTC |
|---|
| BMDB ID | BMDB0000419 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
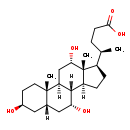
| Common Name | 3b,7a,12a-Trihydroxy-5b-cholanoic acid |
|---|
| Description | 3b,7a,12a-Trihydroxy-5b-cholanoic acid, also known as 3-epi-cholic acid or 3-epi-cholate, belongs to the class of organic compounds known as trihydroxy bile acids, alcohols and derivatives. These are prenol lipids structurally characterized by a bile acid or alcohol which bears three hydroxyl groups. Based on a literature review a significant number of articles have been published on 3b,7a,12a-Trihydroxy-5b-cholanoic acid. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| (3beta,5beta,7alpha,12alpha)-3,7,12-Trihydroxycholan-24-Oic acid | ChEBI | | 3-Epi-cholic acid | ChEBI | | 3-Epicholic acid | ChEBI | | 3beta-Cholic acid | ChEBI | | Isocholic acid | ChEBI | | (3b,5b,7a,12a)-3,7,12-Trihydroxycholan-24-Oate | Generator | | (3b,5b,7a,12a)-3,7,12-Trihydroxycholan-24-Oic acid | Generator | | (3beta,5beta,7alpha,12alpha)-3,7,12-Trihydroxycholan-24-Oate | Generator | | (3Β,5β,7α,12α)-3,7,12-trihydroxycholan-24-Oate | Generator | | (3Β,5β,7α,12α)-3,7,12-trihydroxycholan-24-Oic acid | Generator | | 3-Epi-cholate | Generator | | 3-Epicholate | Generator | | 3b-Cholate | Generator | | 3b-Cholic acid | Generator | | 3beta-Cholate | Generator | | 3Β-cholate | Generator | | 3Β-cholic acid | Generator | | Isocholate | Generator | | 3b,7a,12a-Trihydroxy-5b-cholanoate | Generator | | 3b,7a,12a-Trihydroxy-5b-cholan-24-Oate | HMDB, Generator | | 3b,7a,12a-Trihydroxy-5b-cholan-24-Oic acid | HMDB, Generator | | 3b,7a,12a-Trihydroxy-5b-cholanate | HMDB | | 3b,7a,12a-Trihydroxy-5b-cholanic acid | HMDB | | 3beta,7alpha,12alpha-Trihydroxy-5beta-cholan-24-Oate | Generator, HMDB | | 3Β,7α,12α-trihydroxy-5β-cholan-24-Oate | Generator, HMDB | | 3Β,7α,12α-trihydroxy-5β-cholan-24-Oic acid | Generator, HMDB | | (4R)-4-[(1S,2S,5S,7S,9R,10R,11S,14R,15R,16S)-5,9,16-Trihydroxy-2,15-dimethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-14-yl]pentanoate | Generator, HMDB |
|
|---|
| Chemical Formula | C24H40O5 |
|---|
| Average Molecular Weight | 408.5714 |
|---|
| Monoisotopic Molecular Weight | 408.28757439 |
|---|
| IUPAC Name | (4R)-4-[(1S,2S,5S,7S,9R,10R,11S,14R,15R,16S)-5,9,16-trihydroxy-2,15-dimethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-14-yl]pentanoic acid |
|---|
| Traditional Name | (4R)-4-[(1S,2S,5S,7S,9R,10R,11S,14R,15R,16S)-5,9,16-trihydroxy-2,15-dimethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-14-yl]pentanoic acid |
|---|
| CAS Registry Number | 3338-16-7 |
|---|
| SMILES | [H][C@@]12CC[C@H]([C@H](C)CCC(O)=O)[C@@]1(C)[C@@H](O)C[C@@]1([H])[C@@]2([H])[C@H](O)C[C@]2([H])C[C@@H](O)CC[C@]12C |
|---|
| InChI Identifier | InChI=1S/C24H40O5/c1-13(4-7-21(28)29)16-5-6-17-22-18(12-20(27)24(16,17)3)23(2)9-8-15(25)10-14(23)11-19(22)26/h13-20,22,25-27H,4-12H2,1-3H3,(H,28,29)/t13-,14+,15+,16-,17+,18+,19-,20+,22+,23+,24-/m1/s1 |
|---|
| InChI Key | BHQCQFFYRZLCQQ-UXWVVXDJSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as trihydroxy bile acids, alcohols and derivatives. These are prenol lipids structurally characterized by a bile acid or alcohol which bears three hydroxyl groups. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Steroids and steroid derivatives |
|---|
| Sub Class | Bile acids, alcohols and derivatives |
|---|
| Direct Parent | Trihydroxy bile acids, alcohols and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Trihydroxy bile acid, alcohol, or derivatives
- 3-hydroxysteroid
- 12-hydroxysteroid
- 7-hydroxysteroid
- 3-beta-hydroxysteroid
- Hydroxysteroid
- Cyclic alcohol
- Secondary alcohol
- Carboxylic acid derivative
- Carboxylic acid
- Polyol
- Monocarboxylic acid or derivatives
- Organic oxide
- Alcohol
- Organic oxygen compound
- Hydrocarbon derivative
- Carbonyl group
- Organooxygen compound
- Aliphatic homopolycyclic compound
|
|---|
| Molecular Framework | Aliphatic homopolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Status | Expected but not Quantified |
|---|
| Origin | |
|---|
| Biofunction | Not Available |
|---|
| Application | Not Available |
|---|
| Cellular locations | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-03ec-0439000000-abd2a5c162e23ea0d78f | View in MoNA |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (4 TMS) - 70eV, Positive | splash10-001i-1100049000-86faaffe1dc0eadf9e75 | View in MoNA |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_3) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_4) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_1) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_2) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_3) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_4) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_5) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_6) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_3_1) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_3_2) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_3_3) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_3_4) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_3) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_4) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_2_1) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_2_2) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_2_3) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_2_4) - 70eV, Positive | Not Available | View in JSpectraViewer |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - 40V, Negative | splash10-0a4i-0001900000-ae364a047d832b6cfe45 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0000900000-1566aa316f67cc930ca6 | View in MoNA |
|---|
| LC-MS/MS | LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-0000900000-511e7dba102028a6c2b5 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00dl-0009000000-f5b70cbc59d5ea066854 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00dl-0009000000-b06963261afb6fdb494b | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-02c2-2109000000-87477f9c822aeda6a8e0 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4r-0009700000-8dc05bd9542a54f1f91b | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-052r-0009200000-9a24af3f58b590ac86cd | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4l-9007000000-c0f6af27285559bd789e | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0001900000-450d1e4d25aae2ed99f4 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4r-0005900000-630ea179fce67da6f208 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0r09-0019200000-0754019de78fe2be314d | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0abc-0009400000-fe4f5c75dd7ef6e8d9ce | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-059l-3159100000-f0eff49c7e8da5d00c58 | View in MoNA |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0aor-9640000000-28284141ad8f72ab8fe1 | View in MoNA |
|---|
|
|---|